ELISE: Text-Retrieval by Genetic Programming
The ever-growing size of document collections available over the internet or within intranets of large companies makes accurate and complete human indexation impossible. Automated search systems, able to find documents matching a specified criterion, are the key to make the information stored in these repositories available to the end user.
In these knowledge-extraction systems, designed to retrieve information pertaining to a given topic, the notion of topic is often reduced to a collection of words, and information to a collection of documents. However, word meaning and context, linguistic differences and advertising methods (artificially created relationships between documents, recall of an advertising motto, etc) make the search for singular topics (expressed by rarely used combinations of common words or by words or acronyms with multiple meanings) very disappointing, even with state of the art search engines: relevant information is lost in a sea of off-topic documents.
An additionnal hurdle that faces this task is the difference between meaning, i.e. what a specific user means by a word or a set of words, and sense, i.e. the generally received acception for the same word or set of words. When document repositories contain information in several languages, sometimes mixed in a single document, using distinct abreviation conventions, and covering several disciplines, commercial tools commonly fail to retrieve relevant information, mostly because they need to sacrifice semantic analysis to provide ``efficient'' service and multi-lingual support.
Instead of focusing on improved, efficient generic systems for query analysis and document indexation, we take an orthogonal approach, by putting the emphasis on user-dependant improvement. We aim to improve the retrieval of information relevant to a particular user, through an adaptive learning of the formulations biases and search preferences specific of this user.
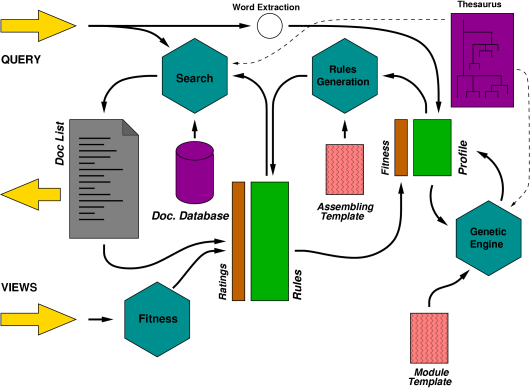
The current version of our adaptive search engine, ELISE (for Evolutionary Learning Interactive Search Engine), relies on a Genetic Programming core to learn a personal profile of expansion rules. Learning is directed by evaluating the quality of the rules, through a measure of the time spent by the user on each returned item.

Much as semantic networks are used in upscale text-retrieval tools to drive a query-expansion step, to enlarge the scope of the query from particular words to actual senses, we use these rules to transform the user's query (a set of words) into a rationalized query to pass to a standard boolean TR tool.
This concept was concretized in a prototype search engine, tailored for pharmaco-medical databases, in collaboration with Novartis Pharma. Tests conduced on references databases (TREC test sets) produced interesting results (see attached bibliography), and real-environment tests are underway.